Loose Leaf: The official blog of American Forests
gary.maguireLoose Leaf is a great blog on tree pests and what can be done to strengthen forests.
Loose Leaf is a great blog on tree pests and what can be done to strengthen forests.
In spring of 2004, WSU initiated a research project to examine factors influencing P. ramorum infection in Christmas tree plantations with funding provided by a two-year USDA Forest Service grant. Work on this project is occurring at the Black Road Christmas Tree Farm near Los Gatos, CA. This is a 23-acre U-cut Christmas tree farm that was established in 1966. Conifers being grown at this site include Douglas-fir, grand fir, giant sequoia, scotch pine, white fir, and California red fir. Some known P. ramorum hosts in the forest adjacent to the edge of the farm include:
California bay laurel, madrone, big leaf maple, toyon, coast redwood, and tanoak. Dieback on some of the grand and Douglas-fir along the interface between the infected forest and the Christmas tree farm appears to have occurred at least 4 years previously, in 2000. During 2005, conditions were very favorable for disease development. Results from this study indicate that most of the infected Christmas trees occurred within 2 meters of the edge of the P. ramorum-infected bay laurel forest canopy. Virtually no infection was evident on Christmas trees that were >5 meters away from the forest edge.
Given recent developments in the UK, where P. ramorum is infecting conifers, further research at WSU-P is being done on the susceptibility of conifer species in western Washington forests.
Symptoms Associated with Inoculation of Stems on Living Douglas-fir and Grand Fir Trees with Phytophthora ramorum 15 KB
Gary Chastagner, Kathy Riley, Katie Coats, Marianne Elliott, Annie DeBauw, and Norm Dart.
In Frankel, Susan J.; Kliejunas, John T.; Palmieri, Katharine M. 2010. Proceedings of the Sudden Oak Death Fourth Science Symposium. Gen. Tech. Rep. PSW-GTR-229. Albany, CA: U.S. Department of Agriculture, Forest Service, Pacific Southwest Research Station. 378 p.
Chastagner, G.A., E.M. Hansen, K.L. Riley, and W. Sutton. 2005. Susceptibility of conifer shoots to infection by Phytophthora ramorum. Sudden Oak Death Science Symposium II, 18-21 January 2005, Monterey, CA.
Chastagner, G.A., K.L. Riley, E.M. Hansen, and W. Sutton. 2004. Christmas tree and conifer nursery stock: Sudden oak death project update. Christmas Tree Lookout 37(3): 10-13.
Chastagner, G.A., E.M. Hansen, K.L. Riley, and W. Sutton. 2004. Susceptibility of conifer shoots to infection by Phytophthora ramorum. Phytopathology 94: S16.
Chastagner, G.A., E.M. Hansen, K.L. Riley, and W. Sutton. 2003. Susceptibility of conifer shoots to infection by Phytophthora ramorum. In: Program and abstract book, sixth international Christmas tree research and extension conference; 2003 Sept. 14-19; Hendersonville, NC; 10-11.
Contact: Gary Chastagner, 253-445-4528 | WSU Puyallup Research & Extension Center, 2606 West Pioneer, Puyallup, WA, 98371-4998 USA
Last updated January 2, 2013
 In southwest Oregon, an aggressive program of cutting and burning host plants in an effort to eradicate Phytophthora ramorum was begun in 2001. It was soon apparent that tanoak (Lithocarpus densiflorus) resprouts were highly susceptible to P. ramorum and that infected sprouts hamper eradication efforts by maintaining inoculum on site.
In southwest Oregon, an aggressive program of cutting and burning host plants in an effort to eradicate Phytophthora ramorum was begun in 2001. It was soon apparent that tanoak (Lithocarpus densiflorus) resprouts were highly susceptible to P. ramorum and that infected sprouts hamper eradication efforts by maintaining inoculum on site.Biological control of tanoak resprouts using the fungus Chondrostereum purpureum. Marianne Elliott, Simon Shamoun, Grace Sumampong, Ellen Goheen, Alan Kanaskie, and Gary Chastagner. Poster presented at 59th WIFDWC, October 2011.
 |
 |
Site # 1002
First bait deployment on March 4, 2010 at Anderson Creek by Jenny Davidson, University of Hawaii |
 |
 |
Location of bait bags, April 1, 2010. Photos by Jenny Davidson. |
| Baiting period | Bait leaf spp. | Phytophthora spp. and others identified |
|
1
|
Rhododendron | P. gonapodyides (4), Trichoderma spp. (1), unknown (5) |
|
1
|
Sword fern (Polystichum munitum) | P. gonapodyides (1), unknown (1) |
|
2
|
Rhododendron | P. gonapodyides (4), Trichoderma spp. (1), fungus (cryptosporiopsis, dermateaceae, pezicula) (1), unknown (2) |
|
2
|
Sword fern | P. gonapodyides (1), unknown (1) |
|
3
|
Rhododendron | P. gonapodyides (3), unknown (5) |
|
3
|
Sword fern | Pythium sp. or Py. undulatum (1), unknown (1) |
|
4
|
Rhododendron | P. gonapodyides (6), unknown (5) |
|
4
|
Dandelion (Taraxacum officinale) | Trichoderma spp, Fusarium spp. |
|
5
|
Rhododendron | P. gonapodyides (8), unknown (4) |
|
5
|
Horsetail (Equisetum spp) | unknown (4) |
|
6
|
Rhododendron | P. gonapodyides (5), unknown (8) |
|
6
|
Red alder (Alnus rubra) | P. gonapodyides (1), unknown (2) |
Mainstem Nooksack river – information about the watershed Whatcom Salmon Recovery 2003
Since 2003, the WSU Puyallup Ornamental Plant Pathology Program has been conducting research on Phytophthora ramorum relating to host range, epidemiology, and evaluating potential control options.
In response to industry concerns, WSU built a quarter-million-dollar biocontainment facility at Puyallup. This facility will greatly increase the capacity of WSU to address critical research questions relating to the establishment, host susceptibility, spread and management of P. ramorum. A molecular lab has also been established to enhance research relating to the detection, spread, and genetics of this pathogen.
Management of P. ramorum
Host testing
Nursery research
Biology of P. ramorum – wild type and non-wild type isolates
In January 2010, Steve Oak (US Forest Svc, Asheville, NC) and Jaesoon Hwang (Clemson University, Clemson, SC) visited western Washington to collect samples from streams known to contain P. ramorum and test out the BOB method, filtration, and leaf baiting for detecting P. ramorum. Dan Omdal (WADNR) and Marianne Elliott (WSU) took them to stream sites where leaf baits were deployed and assisted with collecting samples.
The BOB method will be very useful in situations where it is difficult to return to the site to collect more samples or where the water level in the stream is too low to use a bait bag.
See more photos here.
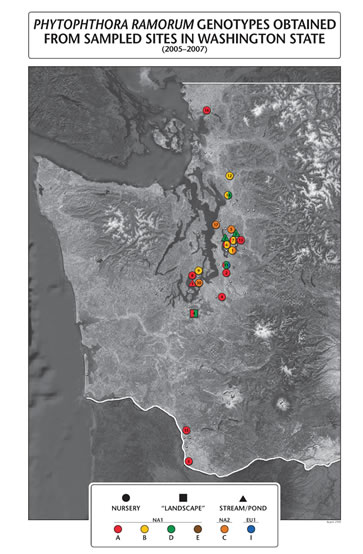 During the past four years (2005-2008), Phytophthora ramorum has been detected in 27 western Washington nurseries. To date, 314 P. ramorum isolates have been collected from the 27 nurseries, as well as two streams, a retention pond, and one landscape site, and genotyped using four microsatellite markers (PrMs5, Pr9C3, PrMs39 and PrMs45).
During the past four years (2005-2008), Phytophthora ramorum has been detected in 27 western Washington nurseries. To date, 314 P. ramorum isolates have been collected from the 27 nurseries, as well as two streams, a retention pond, and one landscape site, and genotyped using four microsatellite markers (PrMs5, Pr9C3, PrMs39 and PrMs45).
All three previously described lineages (EU1, NA1, and NA2) were detected in each of the four years. In this population, the EU1 lineage is represented by one genotype, the NA2 lineage is represented by one genotype, and the NA1 lineage is represented by four genotypes. The NA1 lineage was the most common, occurring in 24 nurseries, one landscape situation, and both streams. The NA2 lineage was detected at 10 nurseries and one stream, while the EU1 lineage was detected at five nurseries. The occurrence of the EU1 lineage in Washington has increased in frequency over the past four years while the overall number of P. ramorum sites and isolates has declined. In four of the five sites where the EU1 lineage occurred, the NA1 lineage was also detected at the same time. At one nursery in 2007, these two lineages were isolated from different branches on the same rhododendron plant. At another nursery in 2008, they were isolated from the same soil bait. Although no genotype detected to date possesses a hybrid of alleles from both the European (EU1) and North American (NA1 and NA2) lineages, the presence of the EU1 and NA1 lineages on the same plant and soil bait illustrates the need for a better overall understanding of the population structure of P. ramorum in nurseries to reduce the risk of hybridized species occurring as a result of sexual recombination.
The APHIS confirmed nursery protocol (CNP) appears to be effective in eradicating the pathogen from infected nurseries in some instances. Fifteen of the 27 positive nurseries were negative for P. ramorum in subsequent years after completing the CNP. On the other hand, 12 nurseries were repeatedly positive for two or more years in a row. At only one repeat-positive nursery did a new genotype appear in the second positive year that was not present in the nursery during the first positive year. Thus it is unclear if inoculum was persisting from year to year or if the same genotypes that were initially detected at 11 of the repeat nurseries was reintroduced in subsequent years. At the six nurseries showing the most genetic variability, three different genotypes were detected in a single year.
from the February 2009 COMTF newsletter
 Western Washington is a “high risk” area for diseases caused by P. ramorum and other Phytophthora species because of favorable environmental conditions and the abundance of susceptible host plants in wildland and urban areas. At present, P. ramorum has only been detected in or near nurseries in western Washington.
Western Washington is a “high risk” area for diseases caused by P. ramorum and other Phytophthora species because of favorable environmental conditions and the abundance of susceptible host plants in wildland and urban areas. At present, P. ramorum has only been detected in or near nurseries in western Washington.
The spread of P. ramorum into the landscape will trigger a series of quarantines that will have a significant economic impact on the horticulture and forest products industry. Stream monitoring programs have been shown to be an effective approach to detect the spread of P. ramorum and focus eradication efforts to high risk areas, thus reducing the threat this pathogen poses to our landscape and forest ecosystems.
SOD National Detection Survey (US Forest Service)
SOD monitoring in California (COMTF site)
P. ramorum surveys in the UK (Forestry Commission)
Stream monitoring for Phytophthora in Washington State, USA: a citizen science project. Presentation at IUFRO 125th Anniversary Congress, Freiburg, Germany Sept. 18-22, 2017.
Community-based stream monitoring for invasive Phytophthoras in western Washington. Marianne Elliott, Gary Chastagner, Katie Coats, Annie DeBauw, and Kathy Riley. Poster, March 2011.
P. ramorum detection and monitoring in western Washington waterways, 2010. Daniel Omdal and Amy Ramsey-Kroll, WA DNR. Poster, Dec. 2010.
The BOB method for detecting P. ramorum in water
Contact: Gary Chastagner, 253-445-4528 | WSU Puyallup Research & Extension Center, 2606 West Pioneer, Puyallup, WA, 98371-4998 USA
Last updated January 2, 2013
Each site had two bait bags containing 10 leaves/sampling period. Bait leaves used were Rhododendron ‘Nova Zembla’ and a ‘wild card’ species of choice. Leaves were sliced in several locations along the midrib to allow for better colonization by Phytophthora. See individual sites for more detailed results. Of these ten leaves, five leaves were cultured and two isolates per leaf were identified using PCR-RFLP and sequencing if necessary, for a total of 10 samples per baiting period. Phytophthora gonapodiydes was found the most often. Unknowns are being sequenced.
1001 – Clarks Creek Downstream
1002 – Anderson Creek
1003 – Evans Creek
1006 – Dogfish Creek